Ass. Prof. Ricardo Aguas

Contact information
Ricardo Aguas
Associate Professor
MAEMOD
Malaria transmission dynamics and optimizing intervention planning.
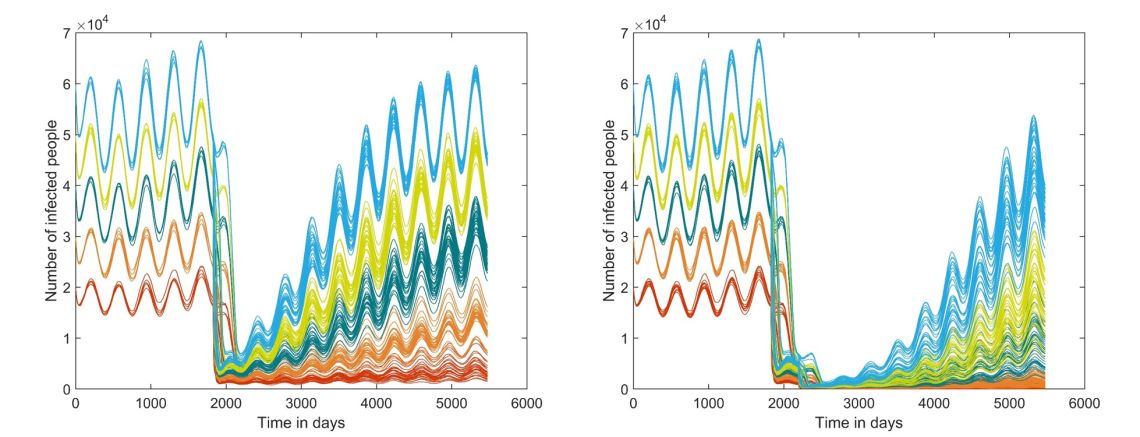
I have been developing a fully dynamic transmission model to simulate targeted malaria elimination strategies (METF) currently undergoing in SE Asia. The model is to be calibrated to the METF transmission settings and run in real time to explore options for study design. It will also be used as a bridge to scale up by projecting the impact of large scale strategy options for larger populations with an additional cost element.
Genotype to phenotype mapping
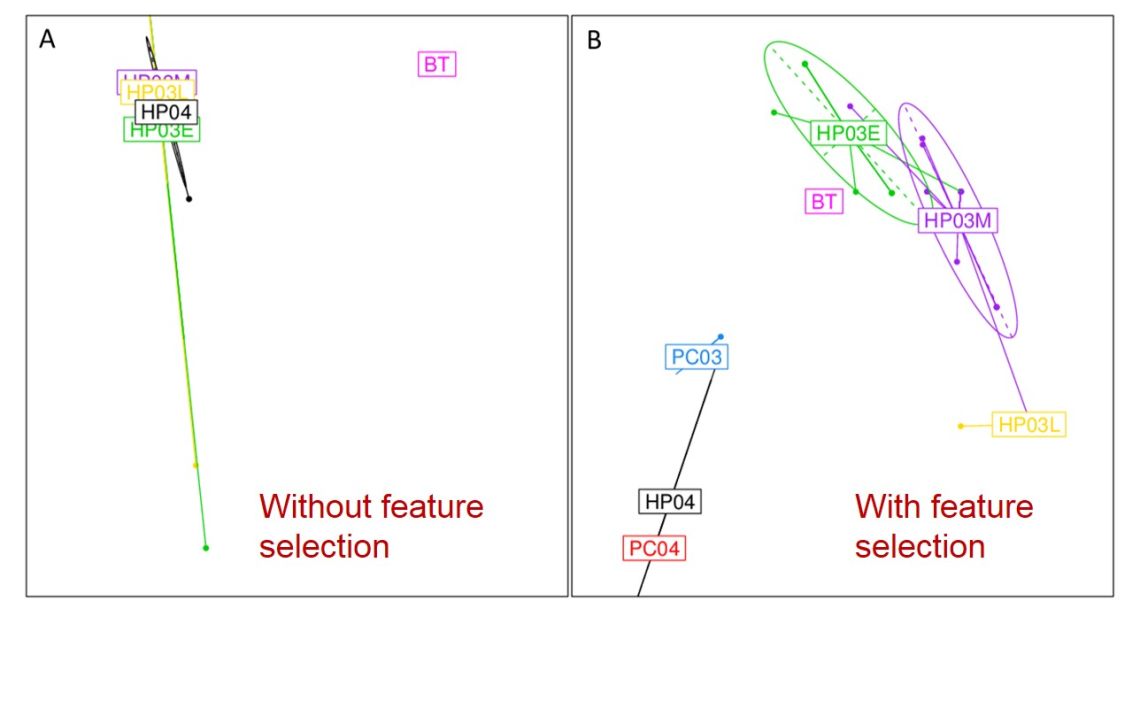
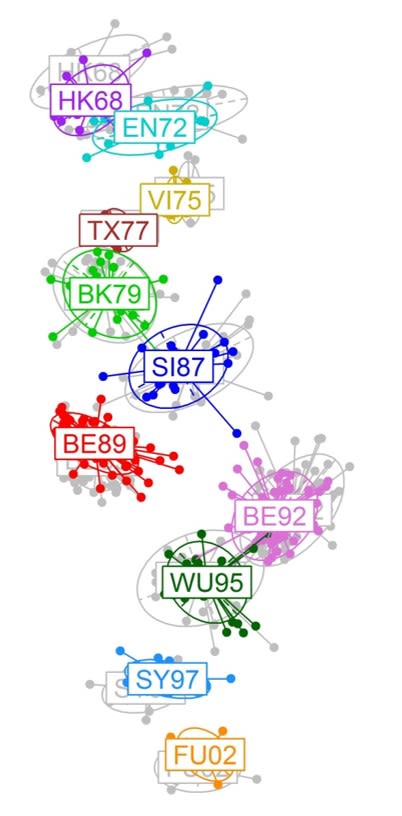
Moving away from genome scan methods used for human GWAS (ultimately inappropriate for the short highly polymorphic genomes of RNA viruses), I have done some work demonstrating the potential of multi-class machine learning algorithms in inferring the functional genetic changes associated with phenotypic change (e.g. a virus crossing the species barrier). These methods allow to uncover a set of features/insights that ultimately could be quite relevant in understanding viral transmission across host species:
- They show that even distantly related viruses within a viral family share highly conserved genetic signatures of host specificity;
- Reinforce how fitness landscapes of host adaptation are shaped by host phylogeny;
- Highlight the evolutionary trajectories of RNA viruses in rapid expansion and under great evolutionary pressure.
These methods can serve as rigorous tools of emergence potential assessment, specifically in scenarios where rapid host classification of newly emerging viruses can be more important than identifying putative functional sites.
Recent publications
-
Understanding COVID-19 testing behaviour in England through a sociodemographic lens: a population based study
Bajaj S. et al, (2024), Lancet Digital Health
-
How to model the impact of vaccines for policymaking when the characteristics are uncertain: A case study in Thailand prior to the vaccine rollout during the COVID-19 pandemic.
Luangasanatip N. et al, (2023), Vaccine, 41, 4854 - 4860
-
SARS-CoV-2 transmission in opposition-controlled Northwest Syria: modelling pandemic responses during political conflict
Marzouk M. et al, (2022), International Journal of Infectious Diseases
-
Estimating the programmatic cost of targeted mass drug administration for malaria in Myanmar
Kyaw SS. et al, (2021), BMC Public Health, 21
-
Algorithm in the Diagnosis of Febrile Illness Using Pathogen-specific Rapid Diagnostic Tests.
Pokharel S. et al, (2020), Clinical infectious diseases : an official publication of the Infectious Diseases Society of America, 70, 2262 - 2269